Respiratory Rate Variability (RRV)#
This example can be referenced by citing the package.
Respiratory Rate Variability (RRV), or variations in respiratory rhythm, are crucial indices of general health and respiratory complications. This example shows how to use NeuroKit to perform RRV analysis.
Download Data and Extract Relevant Signals#
# Load NeuroKit and other useful packages
import matplotlib.pyplot as plt
import pandas as pd
import neurokit2 as nk
In this example, we will download a dataset that contains electrocardiogram, respiratory, and electrodermal activity signals, and extract only the respiratory (RSP) signal.
# Get data
data = pd.read_csv("https://raw.githubusercontent.com/neuropsychology/NeuroKit/master/data/bio_eventrelated_100hz.csv")
rsp = data["RSP"]
nk.signal_plot(rsp, sampling_rate=100) # Visualize
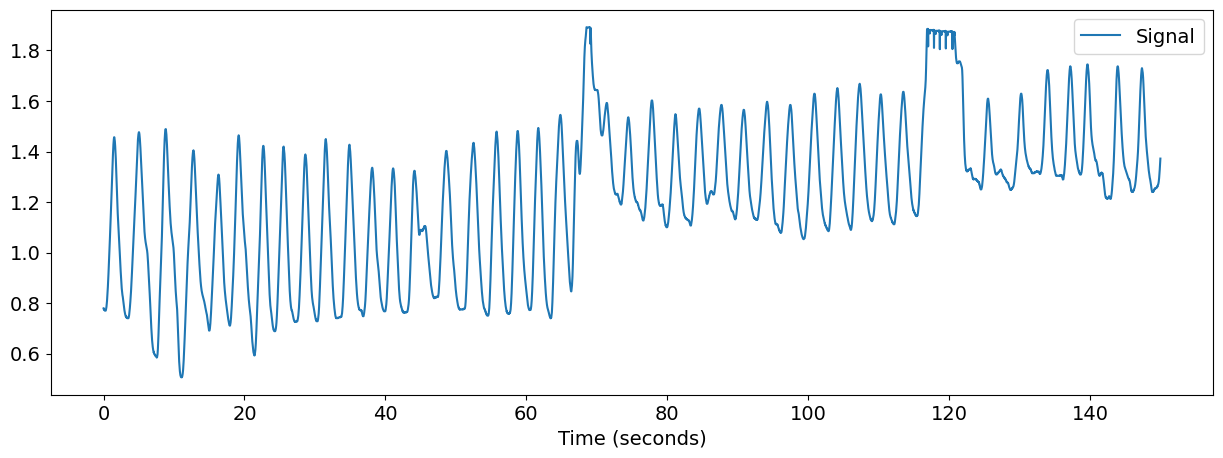
You now have the raw RSP signal in the shape of a vector (i.e., a one-dimensional array). You can then clean it using rsp_clean() and extract the inhalation peaks of the signal using rsp_peaks(). This will output 1) a dataframe indicating the occurrences of inhalation peaks and exhalation troughs (“1” marked in a list of zeros), and 2) a dictionary showing the samples of peaks and troughs.
Note: As the dataset has a frequency of 100Hz, make sure the sampling_rate is also set to 100Hz. It is critical that you specify the correct sampling rate of your signal throughout all the processing functions.
# Clean signal
cleaned = nk.rsp_clean(rsp, sampling_rate=100)
# Extract peaks
df, peaks_dict = nk.rsp_peaks(cleaned)
info = nk.rsp_fixpeaks(peaks_dict)
formatted = nk.signal_formatpeaks(info, desired_length=len(cleaned), peak_indices=info["RSP_Peaks"])
nk.signal_plot(pd.DataFrame({"RSP_Raw": rsp, "RSP_Clean": cleaned}), sampling_rate=100, subplots=True)
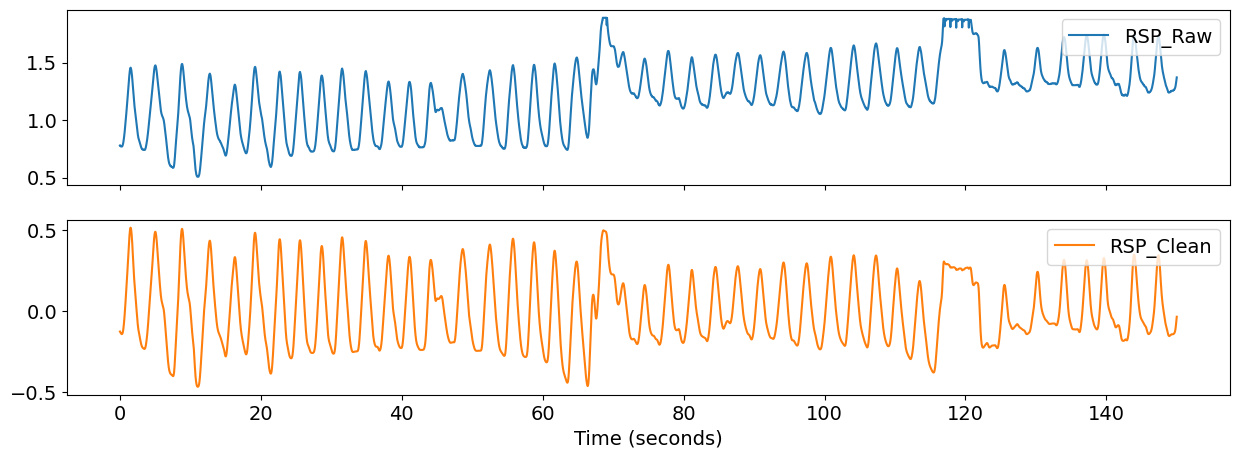
candidate_peaks = nk.events_plot(peaks_dict["RSP_Peaks"], cleaned)
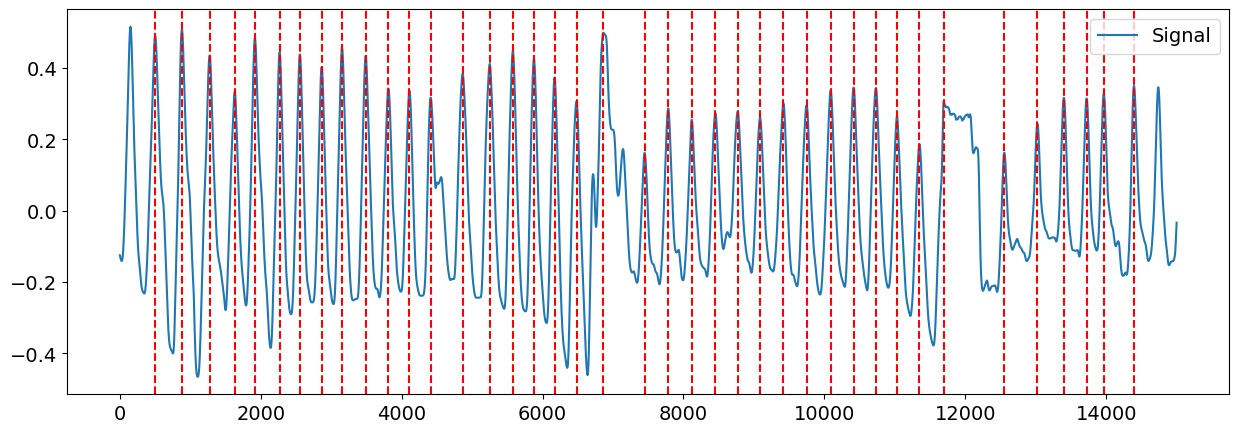
fixed_peaks = nk.events_plot(info["RSP_Peaks"], cleaned)

# Extract rate
rsp_rate = nk.rsp_rate(cleaned, peaks_dict, sampling_rate=100)
# Visualize
nk.signal_plot(rsp_rate, sampling_rate=100)
plt.ylabel("Breaths Per Minute")
Text(0, 0.5, 'Breaths Per Minute')
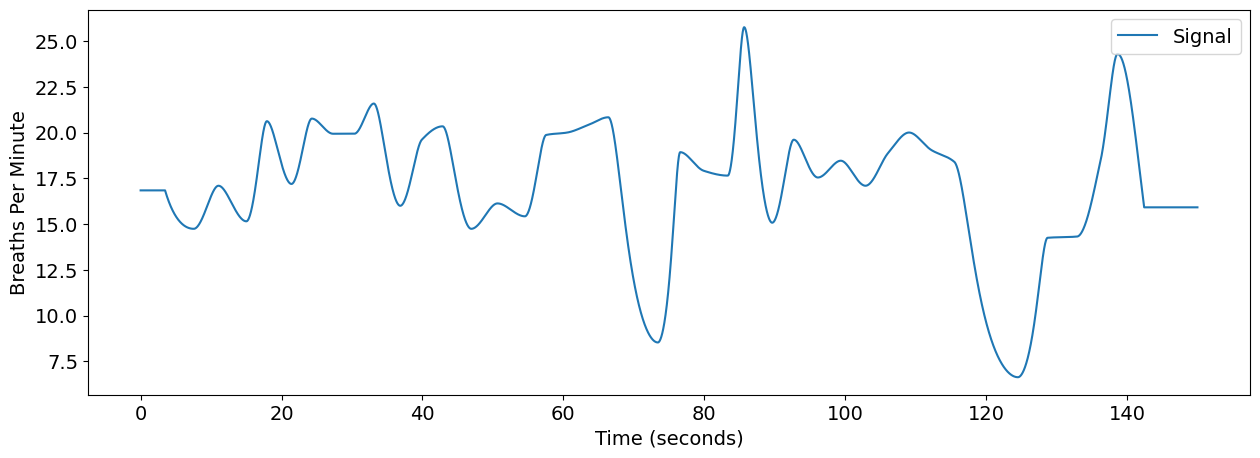
Analyse RRV#
Now that we have extracted the respiratory rate signal and the peaks dictionary, you can then input these into rsp_rrv(). This outputs a variety of RRV indices including time domain, frequency domain, and nonlinear features. Examples of time domain features include RMSSD (root-mean-squared standard deviation) or SDBB (standard deviation of the breath-to-breath intervals). Power spectral analyses (e.g., LF, HF, LFHF) and entropy measures (e.g., sample entropy, SampEn where smaller values indicate that respiratory rate is regular and predictable) are also examples of frequency domain and nonlinear features respectively.
A Poincaré plot is also shown when setting show=True, plotting each breath-to-breath interval against the next successive one. It shows the distribution of successive respiratory rates.
rrv = nk.rsp_rrv(rsp_rate, info, sampling_rate=100, show=True)
rrv
| RRV_RMSSD | RRV_MeanBB | RRV_SDBB | RRV_SDSD | RRV_CVBB | RRV_CVSD | RRV_MedianBB | RRV_MadBB | RRV_MCVBB | RRV_VLF | RRV_LF | RRV_HF | RRV_LFHF | RRV_LFn | RRV_HFn | RRV_SD1 | RRV_SD2 | RRV_SD2SD1 | RRV_ApEn | RRV_SampEn | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1647.714526 | 3562.820513 | 1172.258164 | 1669.813322 | 0.329025 | 0.462475 | 3260.0 | 459.606 | 0.140983 | 0.006583 | 0.02589 | 0.000677 | 38.255223 | 0.781004 | 0.020416 | 1180.736323 | 1163.71824 | 0.985587 | 0.582346 | 1.192138 |
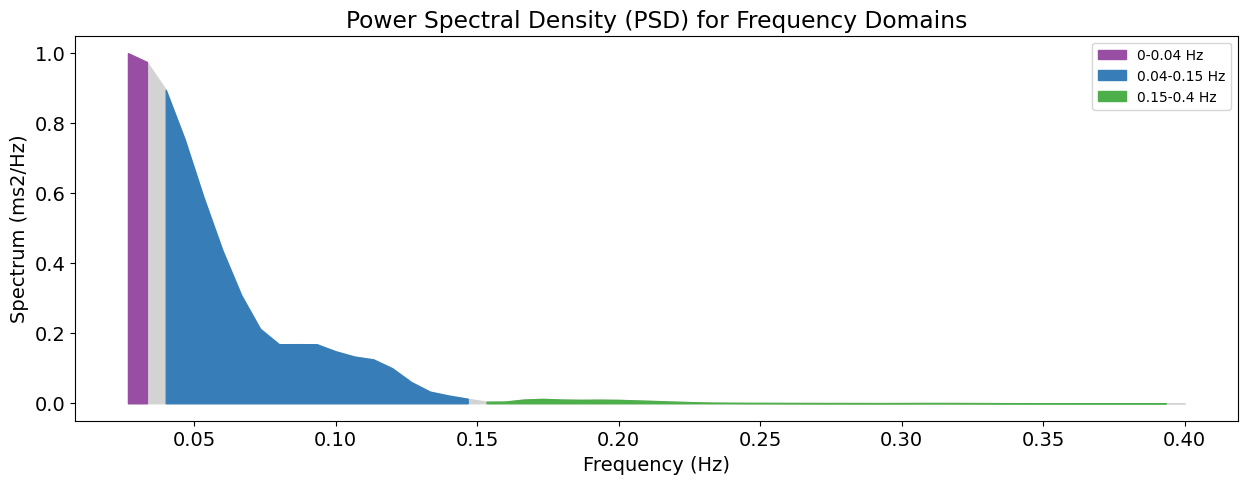
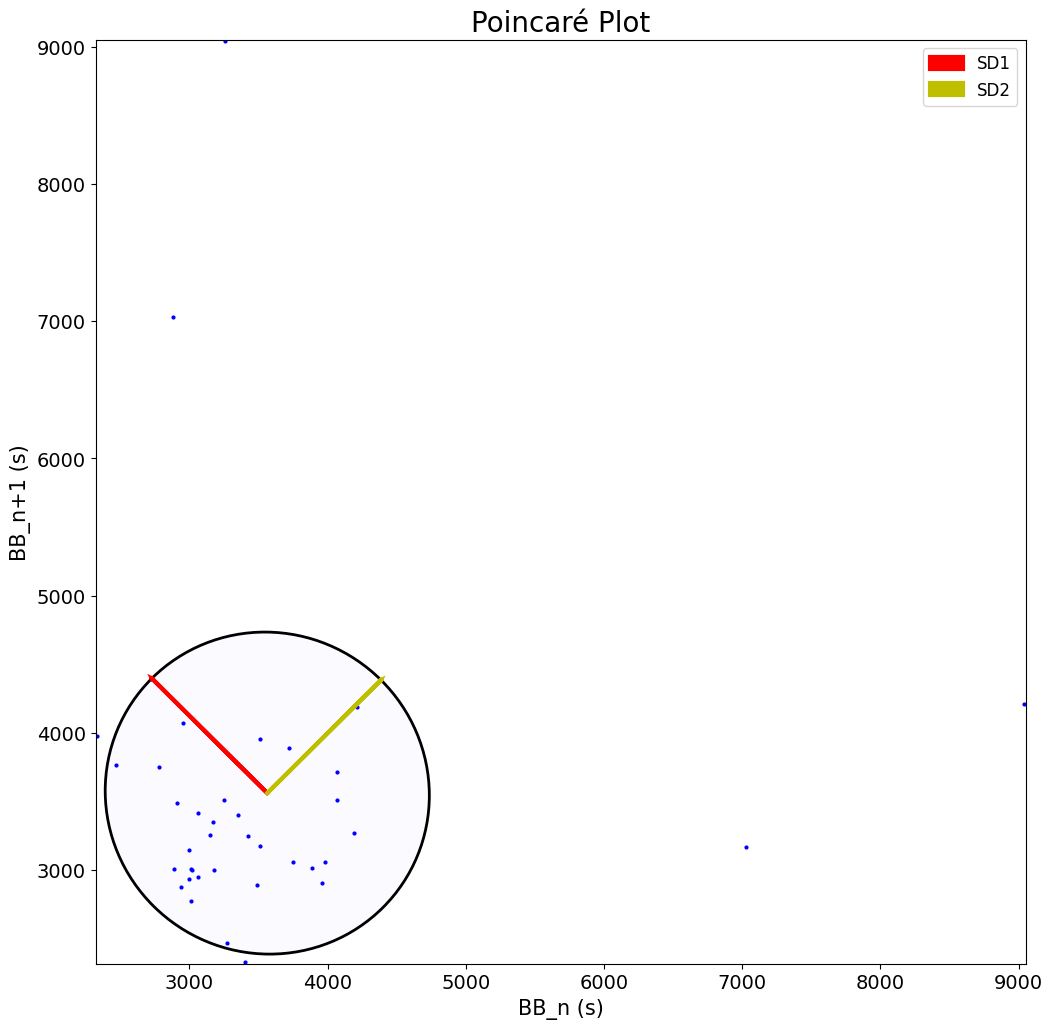
This is a simple visualization tool for short-term (SD1) and long-term variability (SD2) in respiratory rhythm.
